Basic Nipype#
Author: Steffen Bollmann
Date: 17 Oct 2024
Citation and Resources:#
Dataset from OSF#
Shaw, T., & Bollmann, S. (2020). Dataset for Towards Optimising MRI Methods for ChAracterisation of Tissue (TOMCAT) [Data set]. OSF. https://doi.org/10.17605/OSF.IO/BT4EZ
Tools included in this workflow#
Nipype:
Esteban, O., Markiewicz, C. J., Burns, C., Goncalves, M., Jarecka, D., Ziegler, E., Berleant, S., Ellis, D. G., Pinsard, B., Madison, C., Waskom, M., Notter, M. P., Clark, D., Manhães-Savio, A., Clark, D., Jordan, K., Dayan, M., Halchenko, Y. O., Loney, F., … Ghosh, S. (2025). nipy/nipype: 1.8.6 (1.8.6). Zenodo. https://doi.org/10.5281/zenodo.15054147
FSL:
M. Jenkinson, C.F. Beckmann, T.E. Behrens, M.W. Woolrich, S.M. Smith. FSL. NeuroImage, 62:782-90, 2012
Smith S. M. (2002). Fast robust automated brain extraction. Human brain mapping, 17(3), 143–155. https://doi.org/10.1002/hbm.10062
AFNI:
Cox RW (1996). AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput Biomed Res 29(3):162-173. doi:10.1006/cbmr.1996.0014 https://pubmed.ncbi.nlm.nih.gov/8812068/
RW Cox, JS Hyde (1997). Software tools for analysis and visualization of FMRI Data. NMR in Biomedicine, 10: 171-178. https://pubmed.ncbi.nlm.nih.gov/9430344/
SPM12:
Friston, K. J. (2007). Statistical parametric mapping: The analysis of functional brain images (1st ed). Elsevier / Academic Press.
Demonstrating the module system in Python and Nipype#
# we can use module to load fsl in a specific version
import module
await module.load('fsl/6.0.4')
await module.list()
Lmod Warning: MODULEPATH is undefined.
Lmod has detected the following error: The following module(s) are unknown:
"fsl/6.0.4"
Please check the spelling or version number. Also try "module spider ..."
It is also possible your cache file is out-of-date; it may help to try:
$ module --ignore_cache load "fsl/6.0.4"
Also make sure that all modulefiles written in TCL start with the string
#%Module
[]
!bet
/bin/bash: line 1: bet: command not found
Load AFNI and SPM as well#
await module.load('afni/22.3.06')
await module.load('spm12/r7771')
await module.list()
Lmod Warning: MODULEPATH is undefined.
Lmod has detected the following error: The following module(s) are unknown:
"afni/22.3.06"
Please check the spelling or version number. Also try "module spider ..."
It is also possible your cache file is out-of-date; it may help to try:
$ module --ignore_cache load "afni/22.3.06"
Also make sure that all modulefiles written in TCL start with the string
#%Module
Lmod Warning: MODULEPATH is undefined.
Lmod has detected the following error: The following module(s) are unknown:
"spm12/r7771"
Please check the spelling or version number. Also try "module spider ..."
It is also possible your cache file is out-of-date; it may help to try:
$ module --ignore_cache load "spm12/r7771"
Also make sure that all modulefiles written in TCL start with the string
#%Module
[]
Download test data#
%%bash
if [ -f ./sub-01_ses-01_7T_T1w_defaced.nii ]; then
echo "nii Output file exists, not downloading or unpacking again"
else
if [ ! -f ./sub-01_ses-01_7T_T1w_defaced.nii.gz ]; then
echo "nii.gz does not exist. So, it needs to be downloaded."
osfURL="osfstorage/TOMCAT_DIB/sub-01/ses-01_7T/anat/sub-01_ses-01_7T_T1w_defaced.nii.gz"
echo "downloading now ..."
osf -p bt4ez fetch $osfURL ./sub-01_ses-01_7T_T1w_defaced.nii.gz
fi
if [ -f ./sub-01_ses-01_7T_T1w_defaced.nii.gz ]; then
echo "nii.gz exists. So, it needs to be unpacked and deleted"
echo "unpacking now ..."
gunzip ./sub-01_ses-01_7T_T1w_defaced.nii.gz
fi
fi
nii.gz does not exist. So, it needs to be downloaded.
downloading now ...
nii.gz exists. So, it needs to be unpacked and deleted
unpacking now ...
0%| | 0.00/72.7M [00:00<?, ?bytes/s]
0%| | 115k/72.7M [00:00<01:50, 654kbytes/s]
1%| | 639k/72.7M [00:00<00:26, 2.67Mbytes/s]
1%|▏ | 1.02M/72.7M [00:00<00:23, 3.06Mbytes/s]
2%|▏ | 1.39M/72.7M [00:00<00:21, 3.28Mbytes/s]
2%|▏ | 1.77M/72.7M [00:00<00:20, 3.44Mbytes/s]
3%|▎ | 2.16M/72.7M [00:00<00:19, 3.54Mbytes/s]
3%|▎ | 2.54M/72.7M [00:00<00:19, 3.61Mbytes/s]
4%|▍ | 3.01M/72.7M [00:00<00:17, 3.93Mbytes/s]
6%|▌ | 4.05M/72.7M [00:00<00:11, 5.87Mbytes/s]
9%|▊ | 6.36M/72.7M [00:01<00:05, 11.1Mbytes/s]
16%|█▌ | 11.6M/72.7M [00:01<00:02, 23.6Mbytes/s]
20%|██ | 14.7M/72.7M [00:01<00:02, 25.3Mbytes/s]
28%|██▊ | 20.1M/72.7M [00:01<00:01, 33.7Mbytes/s]
35%|███▍ | 25.2M/72.7M [00:01<00:01, 35.4Mbytes/s]
46%|████▌ | 33.1M/72.7M [00:01<00:00, 47.6Mbytes/s]
57%|█████▋ | 41.4M/72.7M [00:01<00:00, 57.1Mbytes/s]
70%|███████ | 51.2M/72.7M [00:01<00:00, 65.8Mbytes/s]
84%|████████▎ | 60.8M/72.7M [00:02<00:00, 65.5Mbytes/s]
100%|██████████| 72.7M/72.7M [00:02<00:00, 34.6Mbytes/s]
%ls
sub-01_ses-01_7T_T1w_defaced.nii
Run nipype pipeline#
%%capture
!pip install nibabel numpy scipy
from nipype.interfaces import fsl
from nipype.interfaces import afni
btr = fsl.BET()
btr.inputs.in_file = './sub-01_ses-01_7T_T1w_defaced.nii'
btr.inputs.frac = 0.4
btr.inputs.out_file = './sub-01_ses-01_7T_T1w_defaced_brain.nii'
res = btr.run()
edge3 = afni.Edge3()
edge3.inputs.in_file = './sub-01_ses-01_7T_T1w_defaced.nii'
edge3.inputs.out_file = './sub-01_ses-01_7T_T1w_defaced_edges.nii'
edge3.inputs.datum = 'byte'
res = edge3.run()
251205-08:20:07,530 nipype.interface WARNING:
FSLOUTPUTTYPE environment variable is not set. Setting FSLOUTPUTTYPE=NIFTI
---------------------------------------------------------------------------
OSError Traceback (most recent call last)
Cell In[7], line 8
6 btr.inputs.frac = 0.4
7 btr.inputs.out_file = './sub-01_ses-01_7T_T1w_defaced_brain.nii'
----> 8 res = btr.run()
10 edge3 = afni.Edge3()
11 edge3.inputs.in_file = './sub-01_ses-01_7T_T1w_defaced.nii'
File /opt/conda/lib/python3.13/site-packages/nipype/interfaces/base/core.py:401, in BaseInterface.run(self, cwd, ignore_exception, **inputs)
399 # Run interface
400 runtime = self._pre_run_hook(runtime)
--> 401 runtime = self._run_interface(runtime)
402 runtime = self._post_run_hook(runtime)
403 # Collect outputs
File /opt/conda/lib/python3.13/site-packages/nipype/interfaces/fsl/preprocess.py:163, in BET._run_interface(self, runtime)
159 def _run_interface(self, runtime):
160 # The returncode is meaningless in BET. So check the output
161 # in stderr and if it's set, then update the returncode
162 # accordingly.
--> 163 runtime = super()._run_interface(runtime)
164 if runtime.stderr:
165 self.raise_exception(runtime)
File /opt/conda/lib/python3.13/site-packages/nipype/interfaces/base/core.py:756, in CommandLine._run_interface(self, runtime, correct_return_codes)
753 cmd_path = which(executable_name, env=runtime.environ)
755 if cmd_path is None:
--> 756 raise OSError(
757 'No command "%s" found on host %s. Please check that the '
758 "corresponding package is installed."
759 % (executable_name, runtime.hostname)
760 )
762 runtime.command_path = cmd_path
763 runtime.dependencies = (
764 get_dependencies(executable_name, runtime.environ)
765 if self._ldd
766 else "<skipped>"
767 )
OSError: No command "bet" found on host 992a9059675a. Please check that the corresponding package is installed.
%ls
AA_Neurodesk_demo_tour.ipynb nipype_short.ipynb
MRIQC.ipynb papermill-slurm-submission-example.ipynb
PyBIDS.ipynb pydra_preproc_ants.ipynb
RISE_slideshow.ipynb sub-01_ses-01_7T_T1w_defaced.nii
bids_conversion.ipynb sub-01_ses-01_7T_T1w_defaced_brain.nii.gz
ds000114/ sub-01_ses-01_7T_T1w_defaced_edges.nii
nipype_full.ipynb
# View 3D data
import matplotlib.pyplot as plt
def view_slices_3d(image_3d, slice_nbr, vmin, vmax, title=''):
# print('Matrix size: {}'.format(image_3d.shape))
fig = plt.figure(figsize=(15, 4))
plt.suptitle(title, fontsize=10)
plt.subplot(131)
plt.imshow(np.take(image_3d, slice_nbr, 2), vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Axial');
plt.subplot(132)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 1),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Coronal');
plt.subplot(133)
image_rot = ndimage.rotate(np.take(image_3d, slice_nbr, 0),90)
plt.imshow(image_rot, vmin=vmin, vmax=vmax, cmap='gray')
plt.title('Sagittal');
cbar=plt.colorbar()
def get_figure():
"""
Returns figure and axis objects to plot on.
"""
fig, ax = plt.subplots(1)
plt.tick_params(top=False, right=False, which='both')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
return fig, ax
import nibabel as nib
from matplotlib import transforms
from scipy import ndimage
import numpy as np
# load data
brain_full = nib.load('./sub-01_ses-01_7T_T1w_defaced.nii').get_fdata()
brain = nib.load('./sub-01_ses-01_7T_T1w_defaced_brain.nii.gz').get_fdata()
edges = nib.load('./sub-01_ses-01_7T_T1w_defaced_edges.nii').get_fdata()
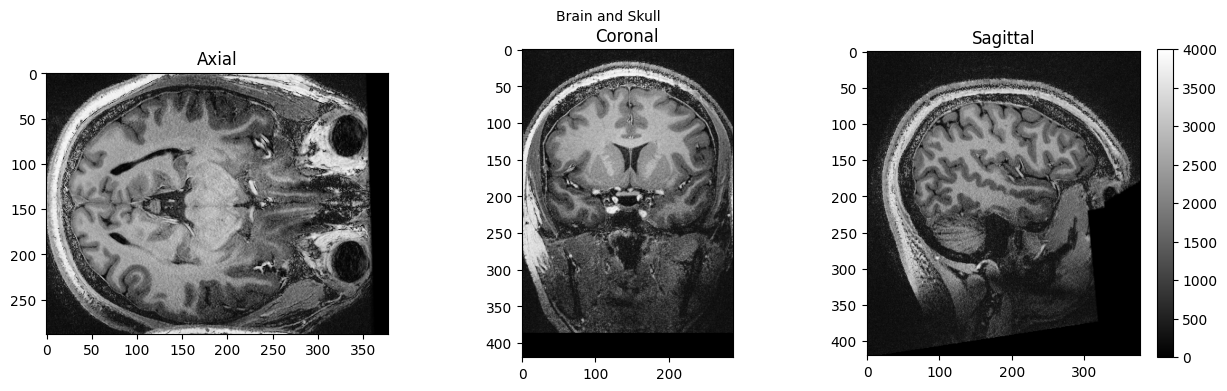
view_slices_3d(brain_full, slice_nbr=230, vmin=0, vmax=4000, title='Brain and Skull')
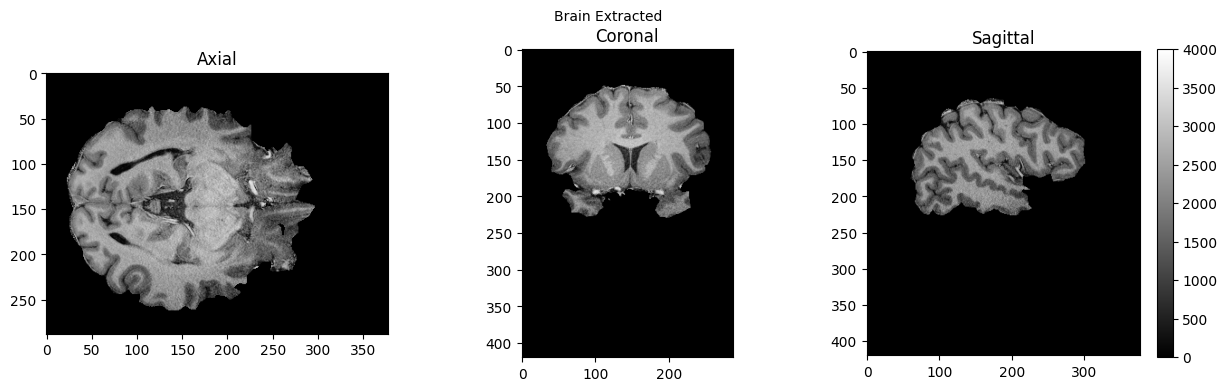
view_slices_3d(brain, slice_nbr=230, vmin=0, vmax=4000, title='Brain Extracted')
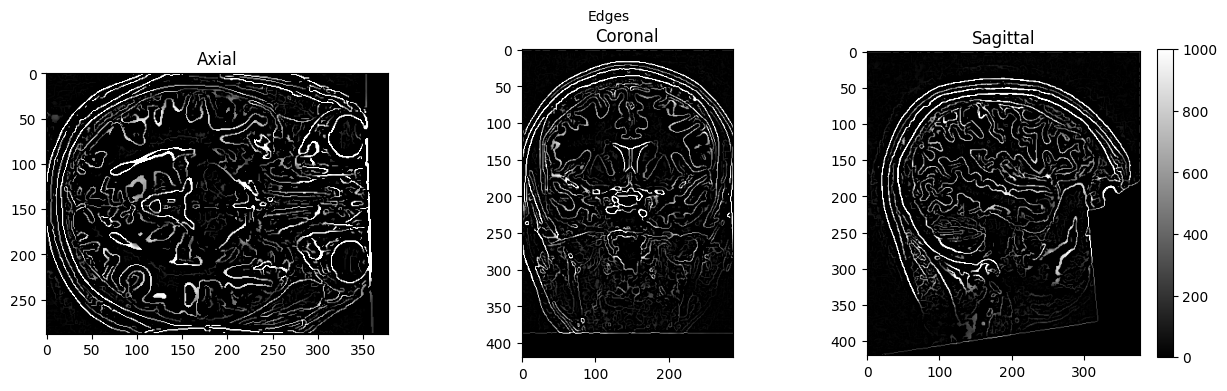
view_slices_3d(edges, slice_nbr=230, vmin=0, vmax=1000, title='Edges')
from ipyniivue import NiiVue
nv = NiiVue()
nv.load_volumes([{"path": "./sub-01_ses-01_7T_T1w_defaced_brain.nii.gz"}])
nv
from IPython.display import Image
Image(url='https://raw.githubusercontent.com/NeuroDesk/example-notebooks/refs/heads/main/books/images/sub-01_ses-01_7T_T1w_defaced_brain.png')

SPM can also be used in such a workflow, but unfortunately, this will trigger a warning “stty: ‘standard input’: Inappropriate ioctl for device”, which you can ignore (or help us to find out where it comes from):
import nipype.interfaces.spm as spm
norm12 = spm.Normalize12()
norm12.inputs.image_to_align = './sub-01_ses-01_7T_T1w_defaced.nii'
norm12.run()
stty: 'standard input': Inappropriate ioctl for device
stty: 'standard input': Inappropriate ioctl for device
<nipype.interfaces.base.support.InterfaceResult at 0x7f7aaa6048d0>
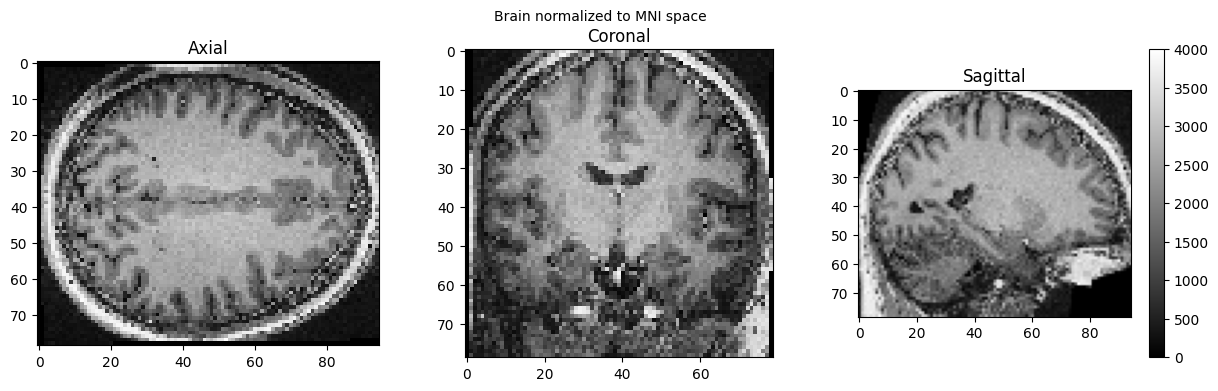
brain_full = nib.load('./wsub-01_ses-01_7T_T1w_defaced.nii').get_fdata()
view_slices_3d(brain_full, slice_nbr=50, vmin=0, vmax=4000, title='Brain normalized to MNI space')
nv = NiiVue()
nv.load_volumes([{"path": "./wsub-01_ses-01_7T_T1w_defaced.nii"}])
nv
Image(url='https://raw.githubusercontent.com/NeuroDesk/example-notebooks/refs/heads/main/books/images/wsub-01_ses-01_7T_T1w_defaced.png')

Dependencies in Jupyter/Python#
Using the package watermark to document system environment and software versions used in this notebook
%load_ext watermark
%watermark
%watermark --iversions
Last updated: 2025-10-31T00:29:05.110304+00:00
Python implementation: CPython
Python version : 3.11.6
IPython version : 8.16.1
Compiler : GCC 12.3.0
OS : Linux
Release : 5.4.0-204-generic
Machine : x86_64
Processor : x86_64
CPU cores : 32
Architecture: 64bit
ipyniivue : 2.3.2
nibabel : 5.2.1
matplotlib: 3.8.4
IPython : 8.16.1
numpy : 2.2.6
nipype : 1.8.6
scipy : 1.13.0
